
DNA and Double D
The latest research on how genetics play a role in developmental dyslexia
By Amielle Moreno PhD.
Gialluisi et al.’s 2021 study, titled Genome-wide association study reveals new insights into the heritability of genetic correlates of developmental dyslexia (DD), is the most ironic study I’ve ever [struggled to] read.
Don’t misunderstand. Misunderstanding is, at least, the first part of my job. It exposes areas where others might also become lost. Throughout my misunderstanding process, it was clear the work conducted by these researchers was undeniably impressive. But, neuroscience research doesn’t get much more ironic than a study on dyslexia that leaves you questioning your own reading ability. I thought I read at a 23rd grade level. All of this made it even more important for me to push through the complex methodology and concepts so you don’t have to.
The following is a short summary of the study and its findings. If you prefer the same information in audio/podcast/comedy form, check out Ep 33 of the Miss Behavior Journal Club, available free of charge for a limited time only.
Intro
Developmental Dyslexia is a learning disorder that affects the ability to read. These specific deficits are not better accounted for by intellectual disabilities or other mental/neurological disorders. Twin studies have reported heritability (h²) of between 40-60%. I was diagnosed with dyslexia as a child and my father likely also had the same disability, on top of being a jerk. Within candidate genes, researchers are looking for the tiny, associated mutations that make a big difference: Single-Nucleotide Polymorphisms (SNPs). To do this, scientists use genome-wide association studies or GWAS. Previously, only two GWAS studies detected criteria meeting associations for genome-wide significance of a SNP or gene of interest for DD (Gialluisi et al., 2019; Truong et al., 2019).
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Nullam eu dignissim tortor, sit amet bibendum lacus. Lorem ipsum dolor sit amet, consectetur adipiscing elit.

DD is associated with brain structure “alterations.” I think, I remember reading about dyslexia and a smaller lateral geniculate nucleus (LGN) about 17 years ago (Livingstone et al., 1991)? But the phenotype likely involves a complex network of brain areas, all fucking up together.
Gialluisi et al. (2021) is a GWAS meta-analysis on developmental dyslexia (DD). By digging through the raw data of multiple studies, the authors set out to determine both what genes and what SNPs are likely suspects in this disorder. They also examined how much DD related SNPs overlap with the SNPs of other psychiatric conditions.
“the authors set out to determine both what genes and what SNPs are likely suspects in this disorder.”
Who’s Asking:
Forty-nine authors, including four last authors, collected data from nine different countries of European ancestry, in six different languages, performing genetic analysis on around two thousand developmental dyslexic cases and more than six thousand control individuals, all to answer the question: “why they don’t read so good?”
Methods:
The genetic association tests and meta-analysis methods section included a lot of words I am not, or am barely familiar with (but that might just be my low reading-based comprehension performance). Terms I had to look up (so you don’t have to) and that I will quickly forget include: minor allele frequency (MAF), Hardy-Weinberg equilibrium (not Harvey Weinstein equilibrium), pairwise genetic distance, and genome-wide heterozygosity values. There was also linkage disequilibrium, copy number variation, variant call rate, and polygenic scores. Half of these, my mechanic said, need to be fixed on my Acura. A crazy level of analysis means a crazy low significance threshold to make up for the fact that there were 18 thousand genes tested (alpha = 2.8 x 10-6).
Gialluisi et 48 peoples, looked at the genome of DD individuals. To create their baseline model, they considered some SNPs can disproportionally contribute to heritability (partitioned heritability). They cited an influential paper in the field of genomic statistics by Finucane et al. (2015).


You see, the genome isn’t a text, in which each sentence is as likely to be read as the last. As I’m sure Finucane would agree, it’s more like Playboy magazine, where some pages are going to become very popular. Also, text isn’t random because each article contains sentences that relate to one another. To explore this analogy further, say there are some different versions of the magazine on the shelves. SNP analysis would be like finding all the sentences in a single magazine that here or there accidentally printed the word ‘boobs’ instead of ‘tits’. These single word polymorphisms would represent SNPs.
No analogy is perfect, Geneticists, so bear with me. Maybe you’re more likely to read some of these ‘tit’ related sentences, based on what is surrounding them, for example, if a sentence is next to a naked-lady picture. There are so many factors that govern if a sentence is going to be read (or a gene is transcribed), including if it’s next to the centerfold (adjacent to a Promoter Upstream Transcription Start Site).
The various analytical approaches utilized in this study take into account the known relationships between groups of SNPs and genes, as they relate to one another in cellular-biological pathways and specific tissues (enrichment analysis). Then the numerous researchers considered histone modifications associated with cells of the central nervous system and corrected or subtracted out categories identified by their baseline model. This allowed them to identify the contributions of central nervous system (CNS) specific variants (stratified linkage disequilibrium regression). Partitioned heritability was used again for genes specifically enriched in major brain regions.
Though I understood a small amount of the statistical corrections necessary here, it was clear the authors were using the most recent and best practices, to ensure accurate representations of the data. Their analysis attempts to take into account what we currently know about the complex nature of genomics.
Findings:
Gialluisi et al. performed association testing at the single-variant level as well as their enrichment analysis to estimate SNP-based heritability.
No single-variant association for people with double Ds reached genome-wide significance. This is different from previous, smaller, non-meta-analysis studies, one of which was published by some of the same authors. This has got to hurt; to wait for a calculation to finish processing and then get a result that doesn’t confirm your previous work. Oof! I guarantee you this particular program script was run again and again and again… with sad but un-biased scientist nearby.

The strongest single-variant association detected was an intronic variant in the gene LOC388780 on chromosome 20. This genomic site is for a non-coding RNA whose function has yet to be determined, but is expressed in the CNS and may play a role in transcriptional regulation. Honestly, all of that means that single gene variations in SNPs might not be the answer for understanding DD and we still don’t know shit.
There was also an enrichment observed at the gene VEPH1 which stands for zone expressed PH domain-containing 1. This protein promotes brain development. They didn’t get any hits on cell-type or brain related histone marker associations.
SNP-based heritability scores indicated about 20-25% of the differences among variance in DD is caused by SNP differences

SNP-based heritability scores indicated about 20-25% of the differences among variance in DD is caused by SNP differences, which is a lot lower than in twin studies suggesting 40-60% heritability. Thus, non-SNP related genetic variants likely contribute to developmental dyslexia. For example, one non-SNP culprit could be copy number variation. This is when the number of copies of a particular gene varies from one individual to the next, or to return to our analogy, Playboy accidently prints the same interview of the actress Partircia Arquett twice in a row.
My first lowlight is one of the multiple reasons this study made me hate myself. I have a bit of experience in genomics but it is not my area of expertise. This article was not written for the population of non-geneticists who will likely be interested in its findings. It took me forever to figure out how SNP-based heritability scores are measured or the various methodological modeling techniques. This paper constantly reminded me I suck at reading.
My second low light brings us to the polygenic score (PGS) findings. A Polygenic score or polygenic risk score reflects an individual’s estimated genetic predisposition for a given trait and is used as a predictor for that trait. The researchers here investigated potential genetic links between dyslexia and other neuropsychiatric disorders.
They used the PGS from other large GWAS studies on neuropsychiatric disorders, as well as intelligence, educational attainment, and cortical brain measures and looked for associations between these scores and their GWAS analysis.
Turns out, I have a lower risk of being an idiot than a lunatic, to borrow some old-school nomenclature. Although the authors ensured direct quotes from their article could not be lifted and interpreted as disparaging of people with DD, this had the unintended effect of making it more difficult to determine the direction of their findings and conclusions.
When I did decipher it, it turns out DD has a significant “risk” of association with fluid intelligence and levels of education. But in what direction? Well, it’s not really a correlation. I don’t intuitively understand Odds Ratios (ORs), which is the way these findings are presented. ORs are notoriously difficult to interpret. You have to report the confidence intervals as well as the p values, which Gialluisi et al. did here. But significance found in an Odds Ratio doesn’t necessarily apply to all members of a population. “There are nearly always a multitude of factors associated with risk” (statisticshowto.com article link below).
The Basics of Interpreting OR:
An odds ratio of more than 1 means there is a higher odds of property B happening with exposure to property A. Example: having prostate cancer increases the odds that you also are a bad driver. What the hell?
OR=1 Exposure does not affect odds of outcome. Example: having prostate cancer is not associated with displaying bad driving. That seems more likely
An odds ratio less than 1 is associated with lower odds. Example: by some freak and unpredictable reason, having prostate cancer makes one want to drive safer and thus there’s a lower risk of bad driving.

Gialluisi et al. report OR for Intelligence is 0.72, with a reasonable CI and very low p value. It’s “significantly associated”. So, intelligence and DD have high odds of association. I went ahead and proved the authors wrong, by having to re-read these results multiple times.
I can’t see how their ‘educational years’ statistic could be applicable, given the young age of most of the subjects. Most of the studies included in the analysis had school-age subjects. Some people with Double Ds return to education as adults, once they are better able to manage their specific difficulties.
DD individuals also had a higher risk of bipolar disorder, ADHD and schizophrenia, with OR scores larger than 1. “BD and SCZ-PGS were associated with an increased risk of DD, explaining 2.8% and 1.4% of its variance, respectively” (Gialluisi et al., 2021). I didn’t find that respectful at all, but it didn’t get me down, perhaps because the OR for major depression was 1.01.
“DD individuals also had a higher risk of bipolar disorder, ADHD and schizophrenia…”
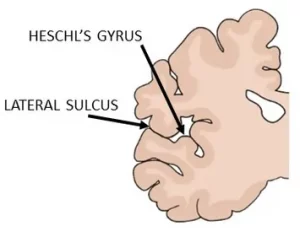
And finally, there’s a protective effect against the risk of Double Ds if you have a thick, hog-sized transverse temporal gyrus. Also known as Heschl’s gyrus, it is located within the primary auditory cortex. It’s important for auditory discrimination and speech perception. Previous studies had not consistently supported the relationship of these star-crossed lovers. But this study is now genetic based evidence for a brain structural feature and dyslexia risk, albeit small and with a number of caveats.
Conclusion:
In general, this study adds to a body of evidence identifying “shared genetic factors influencing educational attainment, general cognition, and more specialized abilities like reading.” Each author contributed copious years of academic training. To think that forty-nine people could all agree on such a complex analytical approach is astounding. They pushed the limits of computational data driven research to produce a paper that you now don’t have to read.
Unformated Bibliography and Links:
Gialluisi A, Andlauer TFM, Mirza-Schreiber N, Moll K, Becker J, Hoffmann P, et al. Genome-wide association study reveals new insights into the heritability and genetic correlates of developmental dyslexia. Mol Psychiatry. 2021 Jul;26(7):3004-3017. doi: 10.1038/s41380-020-00898-x. Epub 2020 Oct 14. PMID: 33057169; PMCID: PMC8505236. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8505236/
Truong DT, Adams AK, Paniagua S, Frijters JC, Boada R, Hill DE, et al. Multivariate genome-wide association study of rapid automatised naming and rapid alternating stimulus in Hispanic American and African–American youth. J Med Genet. 2019. Jmedgenet-2018-105874. https://pubmed.ncbi.nlm.nih.gov/30995994/
Gialluisi A, Andlauer TFM, Mirza-Schreiber N, Moll K, Becker J, Hoffmann P, et al. Genome-wide association scan identifies new variants associated with a cognitive predictor of dyslexia. Transl Psychiatry. 2019;9:77.
How to understand Odds Ratios because you suffer from insomnia or are a masochist:
https://www.statisticshowto.com/probability-and-statistics/probability-main-index/odds-ratio/
Finucane, H., Bulik-Sullivan, B., Gusev, A. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat Genet 47, 1228–1235 (2015). https://doi.org/10.1038/ng.3404
Livingstone, Margaret S., et al. “Physiological and anatomical evidence for a magnocellular defect in developmental dyslexia.” Proceedings of the National Academy of Sciences 88.18 (1991): 7943-7947. https://doi.org/10.1073/pnas.88.18.7943
This is a study from Molecular Psychiatry, titled out of Department of Translational Research in Psychiatry, Max Planck Institute of Psychiatry, the Institute of Human Genetics, University of Bonn, in Germany, Department of Child and Adolescent Psychiatry at Ludwig-Maximilians University, Munich, Germany.The first author is Gialluisi and the last authorS are Schumacher, Nothen, Müller-Myhsok and Körne.

Amielle Moreno PhD.
Scientific Content Creator
Top ,.. top top … post! Keep the good work on !
Good post. I certainly appreciate this website. Stick with it!
Top site ,.. amazaing post ! Just keep the work on !